|
|
MetSite allows users to scan query structures using one of six metal type classifiers. Users must select which classifier they require, additonal site classifiers will be added in future updates to the server.
The MetSite server requires as input a PDB formatted file containing the atomic coordinates of the query protein structure. Please note that MetSite does not require side-chain atoms to be present as the algorithm has been developed for application to low-resolution structures. Therefore only main chain carbon atoms are considered.
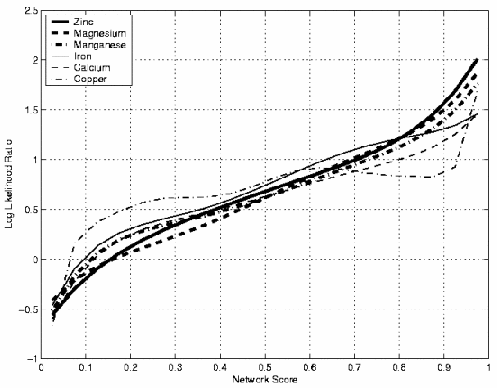
The different site classifers have been optimised to recognise specific site types. To allow outputs from these classifers to be effectivly compared the raw network output is mapped to a log liklihood thereby providing a confidence for the site prediction (Figure 1).
Figure 1: Log-Likelihood

The MetPred program maps the neural network output scores from the selected classifer onto the temperature factor column of the uploaded PDB. Users may then visualise the site prediction using their preferred molecular graphics viewing program.
Please, insert a valid email address to receive the results, which will be returned as soon as they are available. This will usually be a few minutes but may be longer depending on server load. MetSite is free for academic use but access is forbidden for users with commercial email addresses. Enquiries for commercial use should be sent to: dtj@cs.ucl.ac.uk.
The job ID you supply with the structure will be included in the subject line of the email containing the results as an attachment.
Note that, at present, the MetSite server only has limited capacity. We ask that you limit your usage to no more than 20 requests per day, and no more than 5 requests per hour. If you require a larger number of predictions then please get in touch to discuss your requirements.